SONICC®
SONICC®
蛋白质晶体检测
发现隐藏的晶体
发现埋藏在沉淀物中的晶体或肉眼看不到的晶体。
避免错过你感兴趣的晶体
检测薄晶体、小于1 μm 的微晶体和双折射 LCP 中模糊不清的晶体。
聚焦正确的晶体
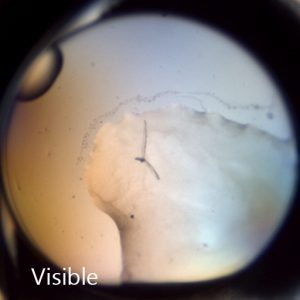
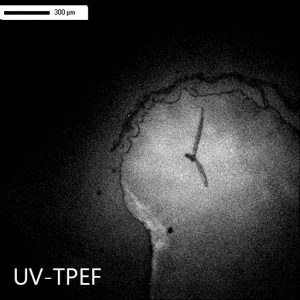
使用集成紫外成像了解您检测到的晶体是否为蛋白晶体。
消除猜测。
SONICC 能够发现通过其他技术无法获得的蛋白质晶体。
非对称晶体的二阶非线性成像 (SONICC) 作为可视化蛋白质晶体的终极成像技术,能够明确地发现隐藏在沉淀物中的晶体或肉眼可见的亚微米晶体。二次谐波 (SHG) 和紫外双光子激发荧光 (UV-TPEF) 作为两种强大的技术,在这个完全自动化的成像系统中集成在一起,可以对用户的高通量结晶板快速成像并有效识别蛋白质晶体。
晶体在纯黑色背景下呈现白色,这样能够帮助用户即使在阴暗的环境下也能识别晶体。SONICC 可以检测极薄的晶体、小于1 μm 的微晶体以及在双折射 LCP 中模糊不清的晶体。
全球主要制药和学术研究实验室一致认为 SONICC是一个非常成功、具有成本效益且运作高效的系统。
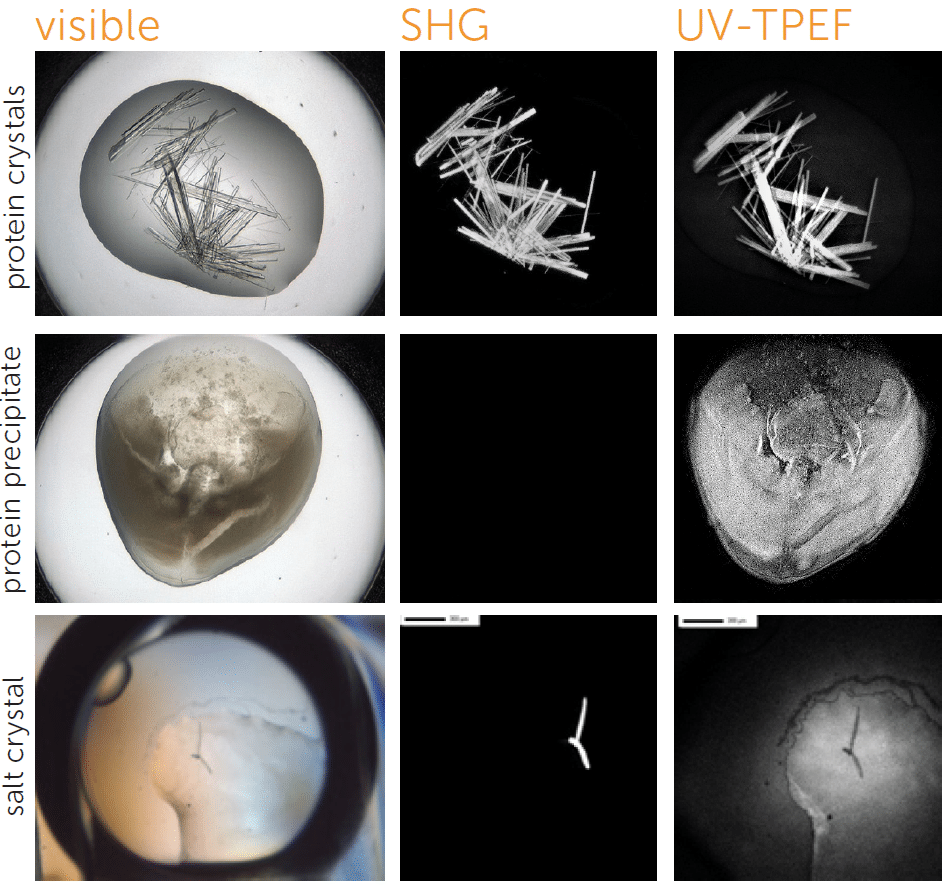
区分盐晶和蛋白质晶体
UV-TPEF(紫外线双光子激发荧光)模式类似于传统的紫外线荧光,并根据紫外线激发的氨基酸(如色氨酸)的荧光创建图像。
UV-TPEF 是一种多光子成像技术,它使用更长的激发波长,有着更好的结晶板兼容性、保证更少的样品损坏以及提供共聚焦成像。
快速识别模糊不清的晶体
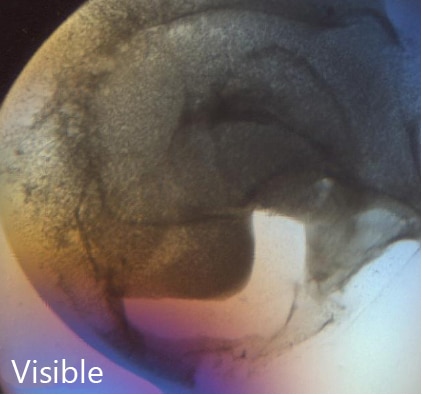
SONICC 技术能够提供高对比度的黑白图像,这样更容易找到隐藏的微晶。下图是在可见光下(左)和 SONICC 在 SHG 模式下(右)对96 孔板的成像 。使用 SHG 成像时,每个含有晶体的结晶板孔都立即显现出来。
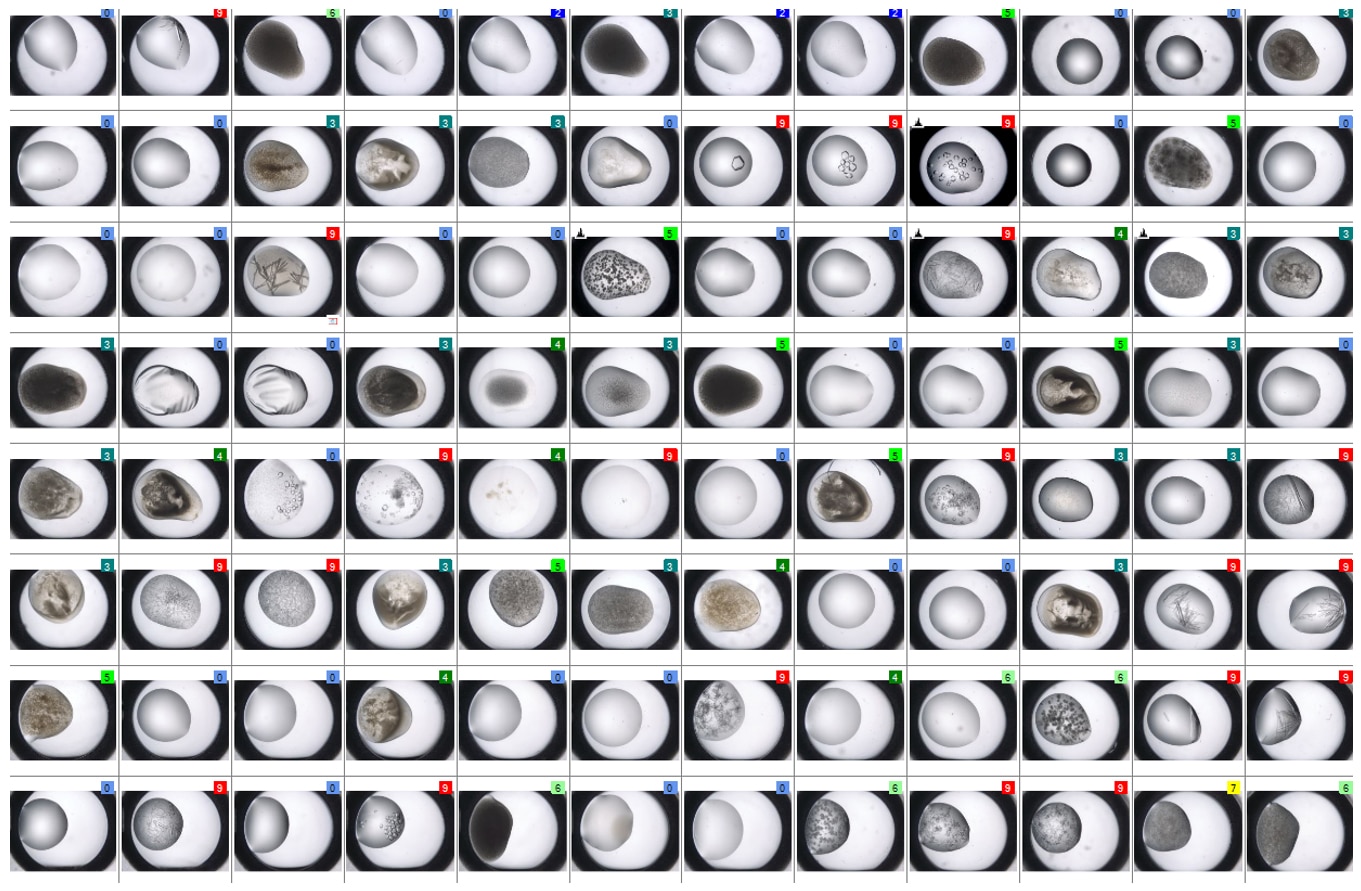

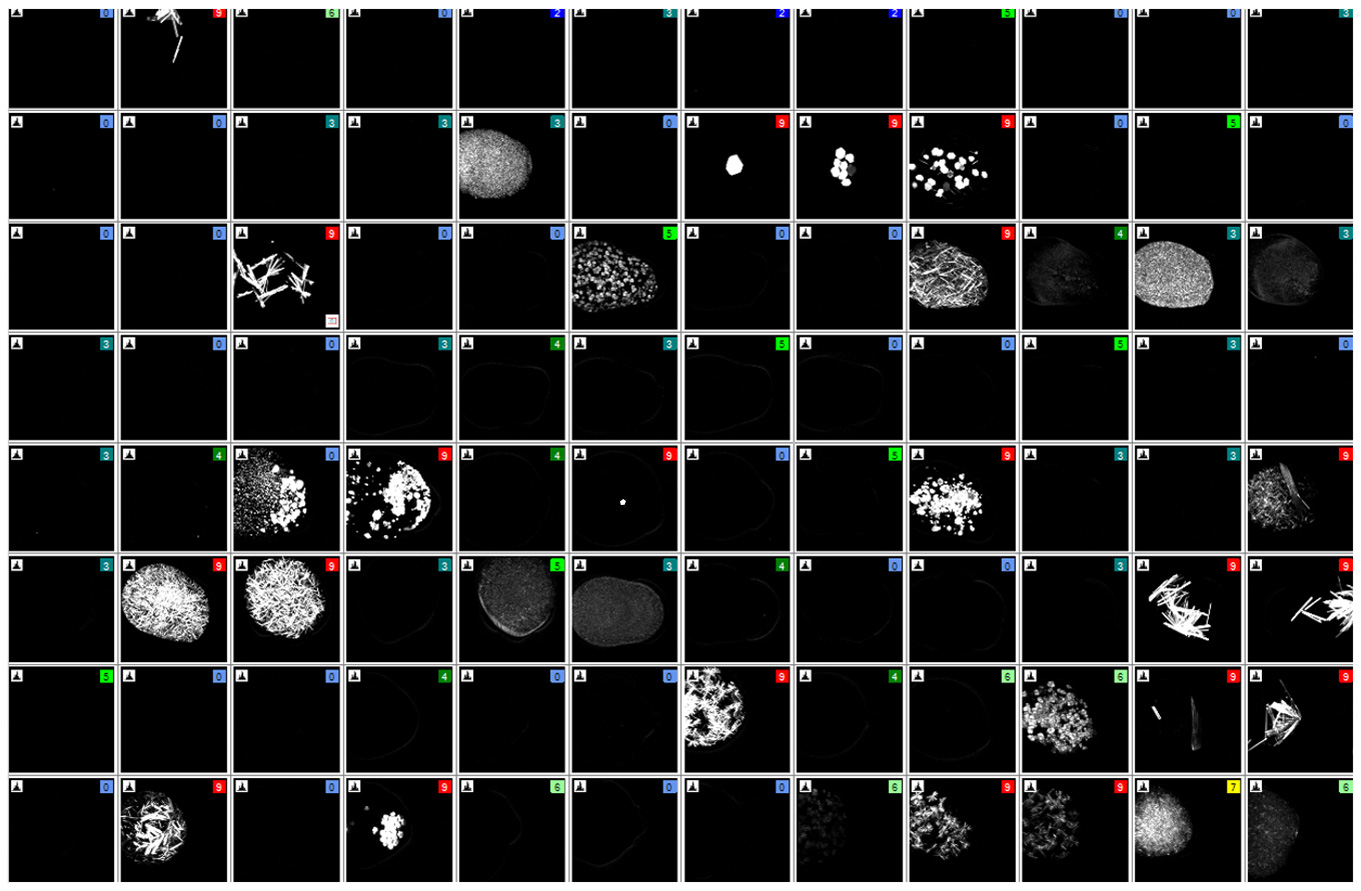
拖动橙色滑块以快速发现使用二次谐波检测到哪些结晶板孔内含有晶体
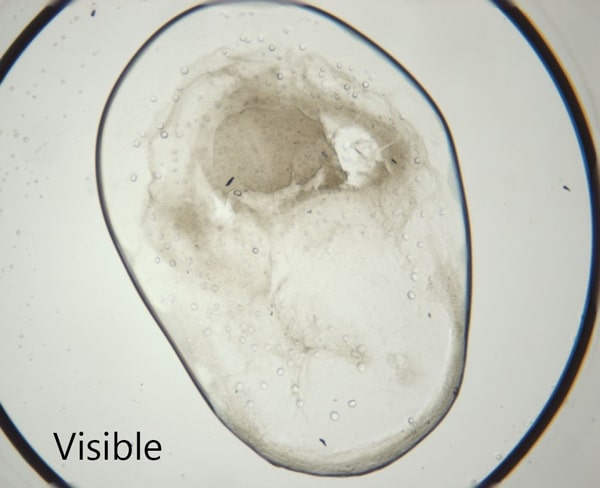
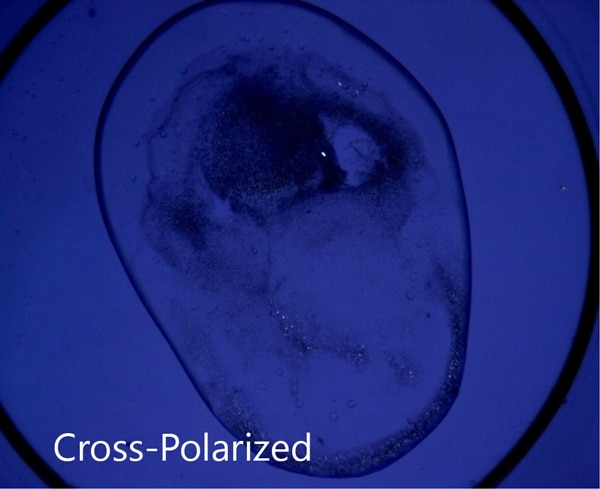
脂质立方相 (LCP) 成像的理想选择
SONICC 极低的检测下限其成为 LCP 结晶板的最佳成像技术。用户使用SONICC可以很容易地对埋藏在混浊的脂质立方相中的小晶体进行可视化成像。液滴定位和自动对焦功能均被用于快速准确地找到 LCP 液滴。同时提供的可见成像技术 - 交叉偏振成像以极快的成像时间给用户提供互补信息。
满足不同用户的工作流程的不同型号
SONICC 提供两种配置以更好地满足用户实验室的需求:
一种是一次只对单块结晶板成像的台式型号。这种最适合对通量无关紧要的实验室。
另一种是内置于 ROCK IMAGER 1000 的集成型号,这种能够让用户装载多达 940 块结晶版并对其自动成像。该型号可以同时对两块结晶板进行成像 - 一块使用可见光成像,一块使用 SONICC成像。这使得用户仅在一天之内便能在对多达 480 块 96 孔板进行可见光成像,以及对多达 96 块 96孔板进行SHG 或 UV-TPEF 成像。

SONICC - 台式:概述
Testimonials

To find microcrystals of membrane protein less than 1 um in LCP, SONICC SHG is really powerful. For this purpose, SHG is much better than UV-TPEF and Cross Polarized in my case.
Kentaro Ihara
RIKEN

It is amazing to see your crystal light up in UV and SHG, for very small crystals it still is a difficult task but with experience, you can resolve a lot of confusion about microcrystals.
Sangita B Patel
Merck

SONICC SHG has been very useful for us in detecting initial crystallization conditions in LCP for membrane protein targets. These conditions usually contain very small crystals and would have been missed by UV imaging especially when the drop is clouded by aggregation or as a result of the opaque LCP matrix.
Kathleen Aertgeerts
Dart Neuroscience, LLC
资源
2020
Towards an Optimal Sample Delivery Method for Serial Crystallography at XFEL
Preventing Bio-Bloopers and XFEL Follies: Best Practices from your Friendly Instrument Staff
Christopher Kupitz and Raymond G. Sierra. Crystals 2020, 10(4), 251; https://doi.org/10.3390/cryst10040251
Received: 5 March 2020 / Revised: 24 March 2020 / Accepted: 24 March 2020 / Published: 27 March 2020
Part of the Methods in Molecular Biology book series (MIMB, volume 2127)
2019
Strategies for sample delivery for femtosecond crystallography
I. Martiel, H. M. Müller-Werkmeister and A. E. Cohen. Acta Cryst. (2019). D75, 160-177 https://doi.org/10.1107/S2059798318017953
Sample Delivery Media for Serial Crystallography
Ki Hyun Nam. Int. J. Mol. Sci. 2019, 20(5), 1094; https://doi.org/10.3390/ijms20051094. Received: 15 February 2019 / Revised: 27 February 2019 / Accepted: 27 February 2019 / Published: 4 March 2019
Successful sample preparation for serial crystallography experiments
John H. Beale, Rachel Bolton, Stephen A. Marshall, Emma V. Beale, Stephen B. Carr, Ali Ebrahim, Tadeo Moreno-Chicano, Michael A. Hough, Jonathan A. R. Worrall, Ivo Tews and Robin L. Owen.
Sample delivery for serial crystallography at free-electron lasers and synchrotrons
M. L. Grünbein and G. Nass Kovacs. Acta Cryst. (2019). D75, 178-191
https://doi.org/10.1107/S205979831801567X
Visualization of protein crystals by high-energy phase-contrast X-ray imaging
M. Polikarpov, G. Bourenkov, I. Snigireva, A. Snigirev, S. Zimmermann, K. Csanko, S. Brockhauser and T. R. Schneider. Acta Cryst. (2019). D75, 947-958
https://doi.org/10.1107/S2059798319011379
Membrane protein megahertz crystallography at the European XFEL
Gisriel, C., Coe, J., Letrun, R. et al. Membrane protein megahertz crystallography at the European XFEL. Nat Commun 10, 5021 (2019) doi:10.1038/s41467-019-12955-3
Pembrolizumab microgravity crystallization experimentation
Reichert, P., Prosise, W., Fischmann, T.O. et al. Pembrolizumab microgravity crystallization experimentation. npj Microgravity 5, 28 (2019) doi:10.1038/s41526-019-0090-3
2018
Lee, D.B., Kim, J., Seok, J.H. et al. Supersaturation-controlled microcrystallization and visualization analysis for serial femtosecond crystallography. Sci Rep 8, 2541 (2018) doi:10.1038/s41598-018-20899-9
2017
Martin-Garcia, J. M., Conrad, C. E., Nelson, G., Stander, N., Zatsepin, N. A., Zook, J., Zhu, L., Geiger, J., Chun, E., Kissick, D., Hilgart, M. C., Ogata, C., Ishchenko, A., Nagaratnam, N., Roy-Chowdhury, S., Coe, J., Subramanian, G., Schaffer, A., James, D., Ketwala, G., Venugopalan, N., Xu, S., Corcoran, S., Ferguson, D., Weierstall, U., Spence, J. C. H., Cherezov, V., Fromme, P., Fischetti, R. F. & Liu, W. (2017). IUCrJ 4, 439-454.
XFELs for structure and dynamics in biology
Spence, J. C. H. (2017). IUCrJ 4, 322-339.
Locating and Visualizing Crystals for X-Ray Diffraction Experiments
Becker M., Kissick D.J., Ogata C.M. (2017) Locating and Visualizing Crystals for X-Ray Diffraction Experiments. In: Wlodawer A., Dauter Z., Jaskolski M. (eds) Protein Crystallography. Methods in Molecular Biology, vol 1607. Humana Press, New York, NY
Structures of riboswitch RNA reaction states by mix-and-inject XFEL serial crystallography
Nature 541, 242–246 (12 January 2017)
doi:10.1038/nature20599
2016
Ishchenko A, Cherezov V, Liu W. Preparation and Delivery of Protein Microcrystals in Lipidic Cubic Phase for Serial Femtosecond Crystallography. Journal of Visualized Experiments : JoVE. 2016;(115):54463. doi:10.3791/54463.
Nonlinear Optical Characterization of Membrane Protein Microcrystals and Nanocrystals
Newman J.A., Simpson G.J. (2016) Nonlinear Optical Characterization of Membrane Protein Microcrystals and Nanocrystals. In: Moraes I. (eds) The Next Generation in Membrane Protein Structure Determination. Advances in Experimental Medicine and Biology, vol 922. Springer, Cham
Dow XY, Dettmar CM, DeWalt EL, et al. Second harmonic generation correlation spectroscopy for characterizing translationally diffusing protein nanocrystals. Acta Crystallographica Section D, Structural Biology. 2016;72(Pt 7):849-859. doi:10.1107/S205979831600841X.
Serial femtosecond crystallography datasets from G protein-coupled receptors
Scientific Data 3, Article number: 160057 (2016)
doi:10.1038/sdata.2016.57
Guiding synchrotron X-ray diffraction by multimodal video-rate protein crystal imaging
Newman JA, Zhang S, Sullivan SZ, et al. Guiding synchrotron X-ray diffraction by multimodal video-rate protein crystal imaging. Journal of Synchrotron Radiation. 2016;23(Pt 4):959-965. doi:10.1107/S1600577516005919.
Transmission electron microscopy for the evaluation and optimization of crystal growth
Stevenson HP, Lin G, Barnes CO, et al. Transmission electron microscopy for the evaluation and optimization of crystal growth. Acta Crystallographica Section D, Structural Biology. 2016;72(Pt 5):603-615. doi:10.1107/S2059798316001546.
A pipeline for structure determination of in vivo-grown crystals using in cellulo diffraction
Boudes M, Garriga D, Fryga A, Caradoc-Davies T, Coulibaly F. A pipeline for structure determination of in vivo-grown crystals using in cellulo diffraction. Acta Crystallographica Section D, Structural Biology. 2016;72(Pt 4):576-585. doi:10.1107/S2059798316002369.
2015
Microfluidic sorting of protein nanocrystals by size for X-ray free-electron laser diffraction
Structural Dynamics 2, 041719 (2015); https://doi.org/10.1063/1.4928688
Serial femtosecond X-ray diffraction of enveloped virus microcrystals
Structural Dynamics 2, 041720 (2015); https://doi.org/10.1063/1.4929410
Batch crystallization of rhodopsin for structural dynamics using an X-ray free-electron laser
Wu W, Nogly P, Rheinberger J, et al. Batch crystallization of rhodopsin for structural dynamics using an X-ray free-electron laser. Acta Crystallographica Section F, Structural Biology Communications. 2015;71(Pt 7):856-860. doi:10.1107/S2053230X15009966.
Serial femtosecond crystallography: the first five years
Schlichting, I. (2015). IUCrJ 2, 246-255.
The detection and subsequent volume optimization of biological nanocrystals
Structural Dynamics 2, 041710 (2015); https://doi.org/10.1063/1.4921199
Caffrey M. A comprehensive review of the lipid cubic phase or in meso method for crystallizing membrane and soluble proteins and complexes. Acta Crystallographica Section F, Structural Biology Communications. 2015;71(Pt 1):3-18. doi:10.1107/S2053230X14026843.
Schubert, R., Meyer, A., Dierks, K., Kapis, S., Reimer, R., Einspahr, H., Perbandt, M. & Betzel, C. (2015). J. Appl. Cryst. 48, 1476-1484.
2014
Crystallization of the large membrane protein complex photosystem I in a microfluidic channel
Abdallah BG, Kupitz C, Fromme P, Ros A. Crystallization of the Large Membrane Protein Complex Photosystem I in a Microfluidic Channel. ACS nano. 2013;7(12):10534-10543. doi:10.1021/nn402515q.
Preparation of microcrystals in lipidic cubic phase for serial femtosecond crystallography
Liu W, Ishchenko A, Cherezov V. Preparation of Microcrystals in Lipidic Cubic Phase for Serial Femtosecond Crystallography. Nature protocols. 2014;9(9):2123-2134. doi:10.1038/nprot.2014.141.
Lee, H.-H., Cherni, I., Yu, H., Fromme, R., Doran, J. D., Grotjohann, I., Mittman, M., Basu, S., Deb, A., Dorner, K., Aquila, A., Barty, A., Boutet, S., Chapman, H. N., Doak, R. B., Hunter, M. S., James, D., Kirian, R. A., Kupitz, C., Lawrence, R. M., Liu, H., Nass, K., Schlichting, I., Schmidt, K. E., Seibert, M. M., Shoeman, R. L., Spence, J. C. H., Stellato, F., Weierstall, U., Williams, G. J., Yoon, C., Wang, D., Zatsepin, N. A., Hogue, B. G., Matoba, N., Fromme, P. & Mor, T. S. (2014). IUCrJ 1, 305-317.
Microseed matrix screening for optimization in protein crystallization: what have we learned?
D’Arcy A, Bergfors T, Cowan-Jacob SW, Marsh M. Microseed matrix screening for optimization in protein crystallization: what have we learned? Acta Crystallographica Section F, Structural Biology Communications. 2014;70(Pt 9):1117-1126. doi:10.1107/S2053230X14015507.
Identifying, studying and making good use of macromolecular crystals
Calero G, Cohen AE, Luft JR, Newman J, Snell EH. Identifying, studying and making good use of macromolecular crystals. Acta Crystallographica Section F, Structural Biology Communications. 2014;70(Pt 8):993-1008. doi:10.1107/S2053230X14016574.
Femtosecond crystallography of membrane proteins in the lipidic cubic phase
Liu W, Wacker D, Wang C, Abola E, Cherezov V. Femtosecond crystallography of membrane proteins in the lipidic cubic phase. Philosophical Transactions of the Royal Society B: Biological Sciences. 2014;369(1647):20130314. doi:10.1098/rstb.2013.0314.
Serial time-resolved crystallography of photosystem II using a femtosecond X-ray laser
Nature 513, 261–265 (11 September 2014)
doi:10.1038/nature13453
2013
Characterization of salt interferences in second-harmonic generation detection of protein crystals
Closser RG, Gualtieri EJ, Newman JA, Simpson GJ. Characterization of salt interferences in second-harmonic generation detection of protein crystals. Journal of Applied Crystallography. 2013;46(Pt 6):1903-1906. doi:10.1107/S0021889813027581.
Madden JT, Toth SJ, Dettmar CM, et al. Integrated nonlinear optical imaging microscope for on-axis crystal detection and centering at a synchrotron beamline. Journal of Synchrotron Radiation. 2013;20(Pt 4):531-540. doi:10.1107/S0909049513007942.
Visualization of membrane protein crystals in lipid cubic phase using X-ray imaging
Warren AJ, Armour W, Axford D, et al. Visualization of membrane protein crystals in lipid cubic phase using X-ray imaging. Acta Crystallographica Section D: Biological Crystallography. 2013;69(Pt 7):1252-1259. doi:10.1107/S0907444913011359.
Imaging of protein crystals with two-photon microscopy
Zhu Q, Toth SJ, Simpson GJ, Hsu H-Y, Taylor LS, Harris MT. Crystallization and Dissolution Behavior of Naproxen/Polyethylene Glycol Solid Dispersions. The journal of physical chemistry B. 2013;117(5):1494-1500. doi:10.1021/jp3106716.
Crystallization and Dissolution Behavior of Naproxen/Polyethylene Glycol Solid Dispersions
Zhu Q, Toth SJ, Simpson GJ, Hsu H-Y, Taylor LS, Harris MT. Crystallization and Dissolution Behavior of Naproxen/Polyethylene Glycol Solid Dispersions. The journal of physical chemistry B. 2013;117(5):1494-1500. doi:10.1021/jp3106716.
2012
DeWalt EL, Begue VJ, Ronau JA, Sullivan SZ, Das C, Simpson GJ. Polarization-resolved second-harmonic generation microscopy as a method to visualize protein-crystal domains. Acta Crystallographica Section D: Biological Crystallography. 2013;69(Pt 1):74-81. doi:10.1107/S0907444912042503.
Modeling the SHG activities of diverse protein crystals
Haupert LM, DeWalt EL, Simpson GJ. Modeling the SHG activities of diverse protein crystals. Acta Crystallographica Section D: Biological Crystallography. 2012;68(Pt 11):1513-1521. doi:10.1107/S0907444912037638.
Femtosecond nanocrystallography using X-Ray Lasers for membrane protein structure determination
Fromme P, Spence JC. Femtosecond nanocrystallography using X-Ray Lasers for membrane protein structure determination. Current opinion in structural biology. 2011;21(4):509-516. doi:10.1016/j.sbi.2011.06.001.
Toth SJ, Madden JT, Taylor LS, Marsac P, Simpson GJ. Selective Imaging of APIs in Powdered Blends with Common Excipients Utilizing TPE-UVF and UV-SONICC. Analytical chemistry. 2012;84(14):5869-5875. doi:10.1021/ac300917t.
2011
Haupert L, Simpson G. Screening of Protein Crystallization Trials by Second Order Nonlinear Optical Imaging of Chiral Crystals (SONICC). Methods (San Diego, Calif). 2011;55(4):379-386. doi:10.1016/j.ymeth.2011.11.003.
Two-photon excited UV fluorescence for protein crystal detection
Madden JT, DeWalt EL, Simpson GJ. Two-photon excited UV fluorescence for protein crystal detection. Acta Crystallographica Section D: Biological Crystallography. 2011;67(Pt 10):839-846. doi:10.1107/S0907444911028253.
Lipidic cubic phase technologies for membrane protein structural studies
Vadim Cherezov, Lipidic cubic phase technologies for membrane protein structural studies, In Current Opinion in Structural Biology, Volume 21, Issue 4, 2011, Pages 559-566, ISSN 0959-440X, https://doi.org/10.1016/j.sbi.2011.06.007.
Detection of Membrane Protein Two-Dimensional Crystals in Living Cells
Gualtieri EJ, Guo F, Kissick DJ, et al. Detection of Membrane Protein Two-Dimensional Crystals in Living Cells. Biophysical Journal. 2011;100(1):207-214. doi:10.1016/j.bpj.2010.10.051.
2010
Nonlinear optical imaging of integral membrane protein crystals in lipidic mesophases.
Kissick DJ, Gualtieri EJ, Simpson GJ, Cherezov V. Nonlinear Optical Imaging of Integral Membrane Protein Crystals in Lipidic Mesophases. Analytical chemistry. 2010;82(2):491-497. doi:10.1021/ac902139w.
2008
Selective Detection of Protein Crystals by Second Harmonic Microscopy
J. Am. Chem. Soc., 2008, 130 (43), pp 14076–14077
DOI: 10.1021/ja805983b
Second Order Non-linear Imaging of Chiral Crystals (SONICC®)
SONICC uses a femtosecond pulsed laser to exploit the frequency-doubling effect found in the majority of protein crystals and produces high-contrast images with negligible background signal.
SONICC has two imaging methods, Second Harmonic Generation (SHG) and Ultraviolet Two-Photon Excited Fluorescence (UV-TPEF). The SHG channel probes crystallinity, and the UV-TPEF channel is specific to proteinaceous samples.
The combined effect of SHG and UV-TPEF imaging is so precise that SONICC can detect microcrystals (<1 um) and distinguish between crystal showers and amorphous aggregates.
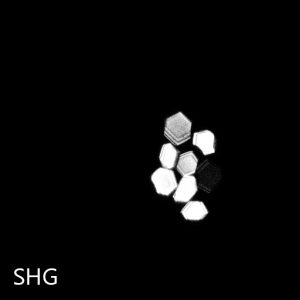
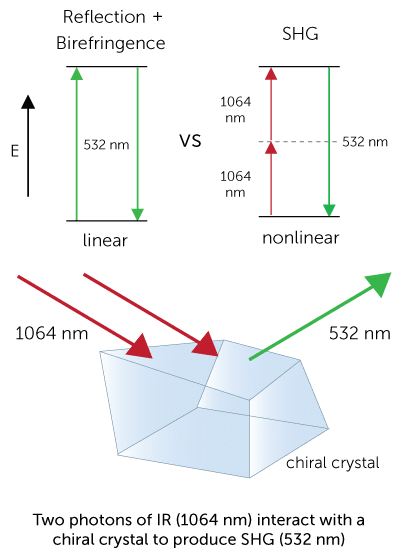
Second Harmonic Generation (SHG)
SHG imaging identifies chiral crystals by imaging the sample with infrared light and detecting the frequency-doubled response, only present in chiral crystals. Chiral crystals are those that lack an internal plane of symmetry. Most salt crystals are symmetric and therefore generate no SHG, whereas all protein crystals are chiral and will generate SHG signal.
Nonlinear effects such as SHG require high electric fields, thus requiring the use of a femtosecond (fs) laser. The laser operates with a pulse width of 200 fs and has high peak powers resulting in nonlinear effects, but the pulses are short enough to prevent sample damage associated with localized heating. Further efforts are taken to prevent sample damage by scanning the laser beam quickly so that it does not remain in one spot long enough to heat the sample.
SHG results from all chiral crystals, including some salts and small molecules that form noncentrosymmetric crystals. These chiral salt and small molecule crystals will result in false positives when imaged in SHG mode. To combat this, SONICC is also equipped with Ultraviolet Two-Photon Excited Fluorescence (UV-TPEF) imaging.
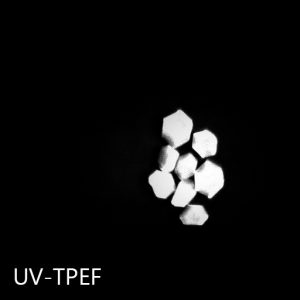
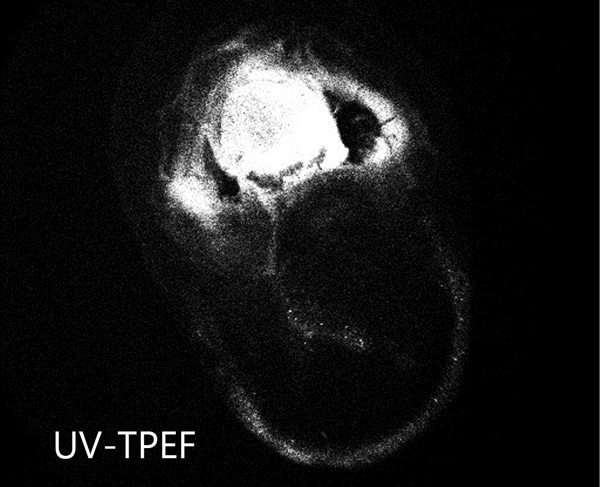
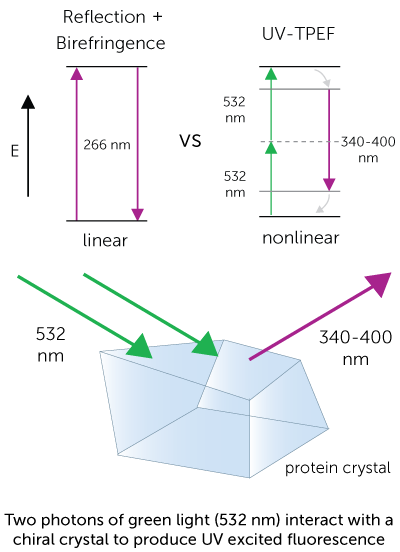
Ultraviolet Two-Photon Excited Fluorescence (UV-TPEF)
UV-TPEF imaging is analogous to traditional UV fluorescence and creates images based on the fluorescence of UV excited amino acids such as tryptophan. UV-TPEF is a multi-photon technique, and therefore uses longer wavelengths of excitation versus traditional linear imaging. This provides significant advantages including compatibility with more plates, less damage to protein and confocal imaging.
In order to probe a sample’s fluorescence, the laser is doubled with a nonlinear optical (NLO) crystal from 1064 nm to 532 nm. The green light (532 nm) is then used to image the sample. The two-photon equivalent of the green light is 266 nm, which excites any tryptophan amino acids that are present in the sample. The two-photon excited fluorescence (350 – 400 nm) is then collected and used to create a fluorescence image.
Imaging Options
| Fixed Zoom | Compound Zoom | Compound Zoom | Compound Zoom | |
|---|---|---|---|---|
| Lens/Objective Options | Asphere 20 mm EFL | Asphere 20 mm EFL | Nikon CFI S Plan Fluor ELWD 20X | 40x Nikon CFI S Plan Fluor ELWD 40X |
| Maximum FOL† | 2.2 x 2.2 mm | 2.2 x 2.2 mm | 1.3 x 1.3 mm | 0.65 x 0.65 mm |
| Lateral Resolution | 4 μm | 4 μm | 2 μm | 1.1 μm |
| Effective N.A.§ | 0.15 | 0.15 | 0.29 | 0.51 |
| Working Distance | 13.5 mm | 13.5 mm | 7.5mm | 3.5 mm |
† Continuous zoom is available by controlling the angle of the scanning mirrors.
§ N.A. (numerical aperture) is proportional to the detection limit of SHG.
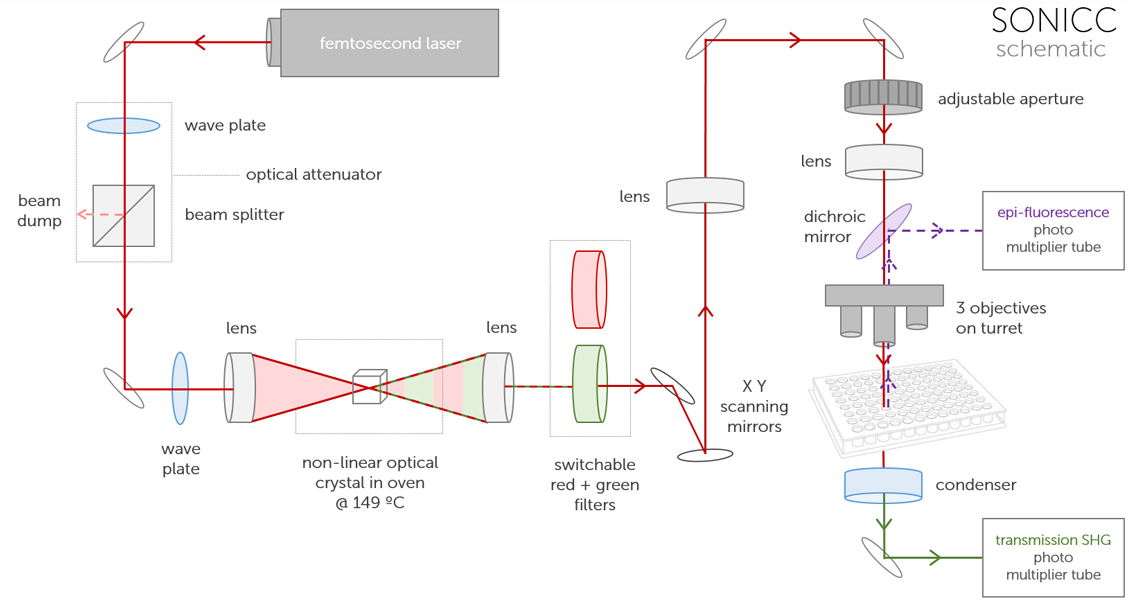
How Does it Work?
1. A femtosecond fiber laser is used to generate the 1064 nm incident IR light.
2. Depending on the mode selected, the wavelength of the laser used to image the sample is changed from IR to green through a nonliner optical crystal.
3. The sample is raster scanned with a resonant mirror along the fast axis and stepped with a galvo-driven mirror for the slow scan axis.
4. A high numerical aperture (NA) objective is used to focus the laser beam onto the sample.
5. The green SHG signal is collected in the transmission direction.
6. UV fluorescence generated in the UV-TPEF mode is collected in the reflected (epi) direction.
7. Photo Multiplier Tubes (PMTs) detect the nonlinear response via photon counting which images are created from.
Background Questions
Are all protein crystals detectable?
Almost all molecules that have a chiral center form a chiral crystal. Therefore, most proteins will form chiral crystals that are detectable via SONICC® (Second Order Non-linear Imaging of Chiral Crystals). Over 99% of the proteins in the Protein Data Bank have a space group that is detectable with SONICC. Crystals with extremely high symmetry classes will generate less SHG signal. Although theoretically all protein crystals should be detected with SHG, that is not always experimentally the case. The higher the space group, the less SHG is generated. Also, the hyperpolarizability (ie the SHG efficiency) of the protein significantly impacts the amount of light generated. Some proteins interact more favorably with light and therefore produce higher SHG signal, where other proteins hold their electrons closely, have a low hyperpolarizability and not give a strong, detectable SHG response.
What is SHG?
SHG stands for Second Harmonic Generation and is a nonlinear optical process. In intense electric fields (i.e. in the presence of a femtosecond laser) the distance between the electrons and the nucleus are distorted (anharmonicity) resulting in non-linear optical effects such as SHG where the frequency of the outgoing light is twice that of the incident (i.e. 1064 nm incident results in 532 nm exiting).
What does “chiral” mean?
A chiral molecule, or in this case a chiral crystal, is a crystal that lacks an internal plane of symmetry, and thus its mirror image is nonsuperimposable. Achiral crystals are symmetric and therefore produce SHG in equal and opposite directions that sum to a net zero signal.
Will salts produce signal?
They can if they are chiral, but the majority of salts are achiral and therefore do not generate SHG signal.
How is SONICC different than fluorescent imaging?
Fluorescent imaging takes advantage of either the endogenous fluorescence of the protein or the use of fluorescent tags. Although fluorescence is bright and easily detectable, it is generated from solubilized and aggregated proteins as well as crystallized proteins. The background from the solubilized protein decreases the signal to noise ratio significantly, resulting in false positives. SONICC, on the other hand, is only sensitive to crystallized proteins.
How does UV-TPEF compare to traditional UV imaging?
Both imaging methods probe the amino acids present in proteins that are excited by UV light (~280 nm). However, with UV-TPEF, the incident wavelength is 532 nm instead of UV and has less energy. UV imaging can cause the breakage of disulfide bonds but by using green light instead to excite, damage does not occur.
With which platforms are SONICC compatible?
SONICC is compatible with all optically-accessible plates and seals.
Will the laser damage my crystals?
Experiments show no detectable damage to protein crystals. In one experiment, a protein crystal was imaged on one half with excessive laser input. X-ray diffraction was obtained from both the exposed and un-exposed halves of the crystal. Both sides diffracted to within expected resolution (~2 Ã) and within statistical variation (i.e. there was no statistical difference between the diffraction of either side). SONICC has also imaged live cells with no observed impact (they remained adhered to a polylysine-coated slide).
Can I use SONICC if my sample is fluorescent?
Yes. As long as fluorescence is Stokes-shifted by 10 nm, the fluorescence will not be detected nor interfere with the SHG signal.
Specification Questions
*Please note that each crystal will generate different intensities of SHG signal depending on size, orientation, space group and quality, as well as the acquisition time and incident intensity.
How small of a crystal can SONICC detect?
Theoretically, the lower limit of detection can be estimated by the forward-to-backward ratio of the SHG signal. Based on the coherence length of the generated SHG signal and the refractive index of the material, this lower limit ranges from 90 nm – 300 nm in thickness. In practice, 1 μm^3 crystals can be routinely detected. 2D crystals have also been routinely imaged with a signal to noise ratio >30.
What is the spatial resolution?
Crystals can be resolved to 2 μm (detection limit is much lower).
What is the laser´s Z-resolution and how deeply can it penetrate?
The laser focuses to a width of ∼50 μm and can image drops >3 mm tall with multiple Z-steps, called “slices”.
How fast is SONICC?
The current electronic package allows 512 x 512 image acquisition for one Z-slice in 500 ms. This corresponds to eight traces of the fast-scanning mirror per line. A one-drop 96-well plate can be imaged with visible light in 3 minutes and with SHG in 15 minutes with eight Z-slices per drop.
Computer Requirements
The standard SONICC controlling computer offered with the instrument supports imaging with continuous multiple focus level imaging (z-slicing) and processing of all images.
SONICC uses the ROCK IMAGER control software which operates with 64-bit Windows 10 or 64/32 bit Windows 7.
Electrical Specifications
SONICC - Benchtop
- Robotics: 100-240 VAC, 50-60 Hz, 600 W max, 1 PH
- UPS (uninterrupted power supply) for robot and laser: 1500 W
- Computer: 100-240 VAC, 50-60 Hz, 525 W max, 1 PH
SONICC - Integrated with ROCK IMAGER 1000 - Dual Imager
- Temperature Regulation: 100-240 V, 50-60 Hz, 750W max, 1 PH
- Robotics: 100-240 V, 50-60 Hz, 270 W max, 1 PH
- Computer: 525 W max, 1 PH
- UPS (uninterrupted power supply) requirement: 4700 W
Physical Dimensions
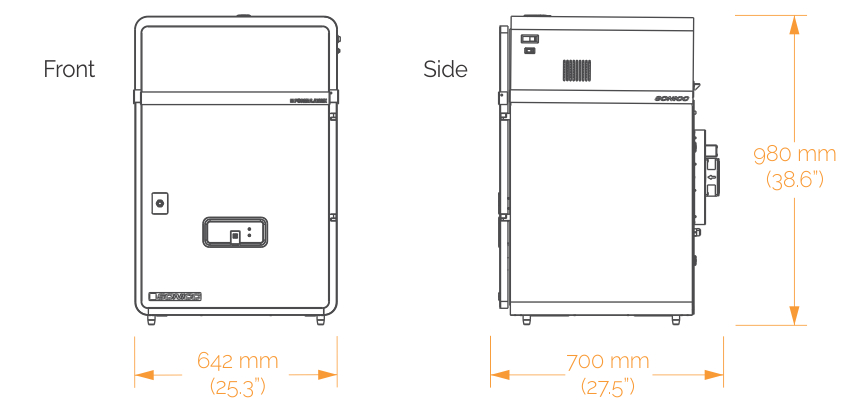
SONICC Benchtop
- Width: 642 mm (25.3")
- Depth: 700 mm (27.5")
- Height: 980 mm (38.6")
- Weight: 85 kg (188 lb)
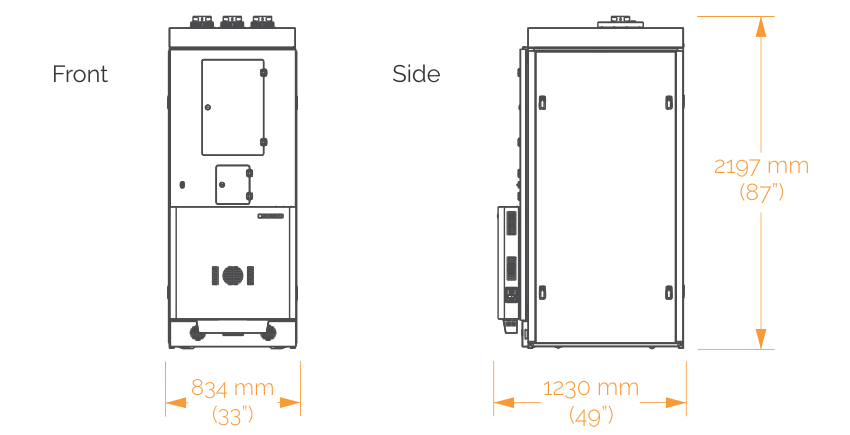
SONICC Integrated Dual Imager
- Width: 834 mm (33")
- Depth: 1230 mm (49")
- Height: 2197 mm (87")
- Weight: 462 kg (1019 lbs)