Understand More About Your Crystals
Expand your understanding of your crystals by imaging in up to three different wavelengths, utilizing both ultraviolet (UV) and visible fluorescence imaging. Determine if your crystals are protein-protein complexes or of a single protein.
Distinguish
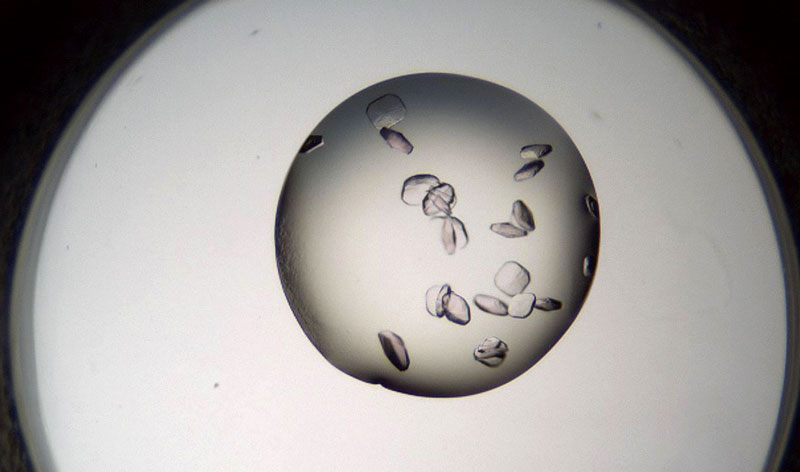
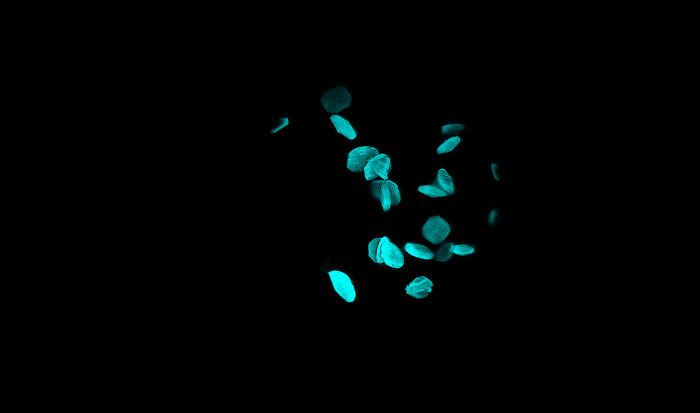
With UV & visible fluorescence imaging, it’s easy to find hidden protein crystals & discriminate them from salt, even protein with little-to-no tryptophan.
Detect Protein-Protein Complexes
Using fluorescent dyes and visible fluorescence imaging, you can differentiate between crystals of a protein-protein complex and crystals of a single protein.
Fast
96-well plates can quickly be imaged in about five minutes using visible fluorescence. UV imaging times vary based on tryptophan concentration but average about 10 minutes per plate.
See Greater Detail
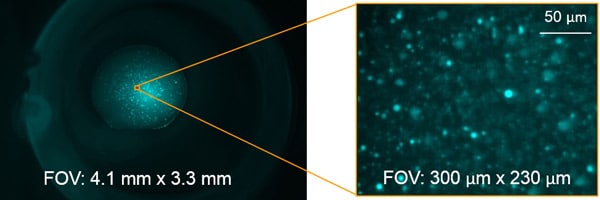
With 3 automated objectives, you can zoom out to see the entire drop or zoom in to see your crystals in greater detail.
Sensitive Crystal Detection
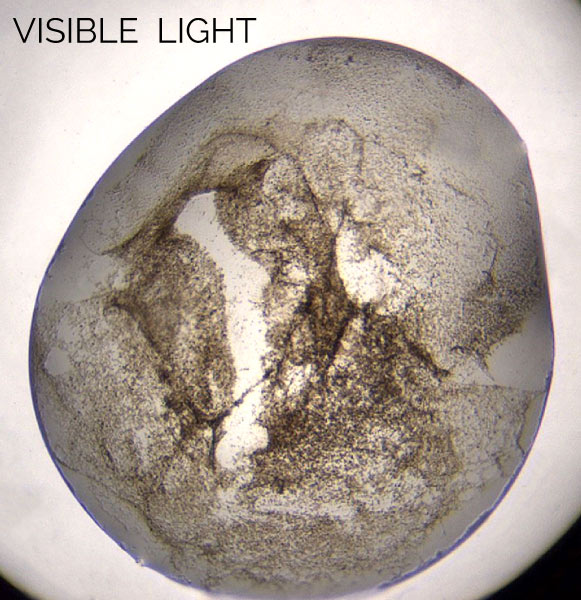
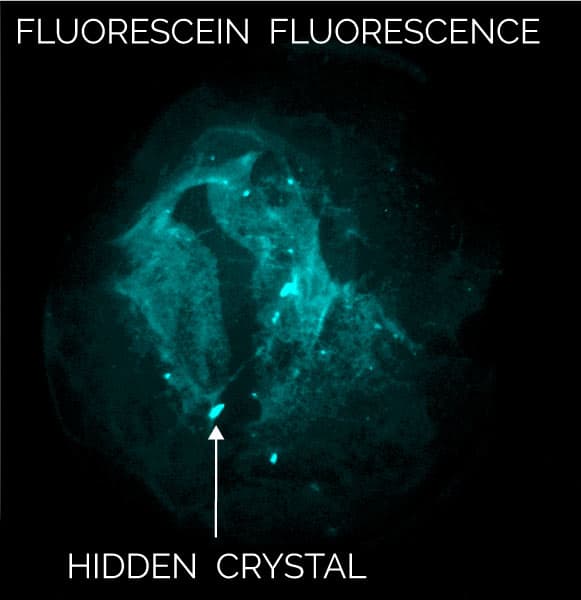
Multi-Fluorescence Imaging (MFI) provides the flexibility to image in 3 different wavelengths of the user’s choosing including UV and visible fluorescence to easily detect protein crystals, even those buried under precipitate.
With UV imaging, drops are illuminated with UV light and the fluorescence generated from aromatic amino acids like tryptophan are detected to create an image.
By labeling only 0.1% of a sample with a fluorophore, visible fluorescence imaging provides the ability to detect proteins with little-to-no tryptophan. This technique acquires high contrast images in very short imaging times with no known sample damage.
Find Protein-Protein Complexes
With MFI, you can now differentiate between crystals of a protein-protein complex and crystals of just one protein. Simply, label the two proteins, or subunits, prior to crystallization with two different amine-reactive dyes and then image your drop at the two corresponding wavelengths. Crystals that fluoresce at both wavelengths are protein-protein complexes, whereas those that fluoresce at only one wavelength are of a single protein.
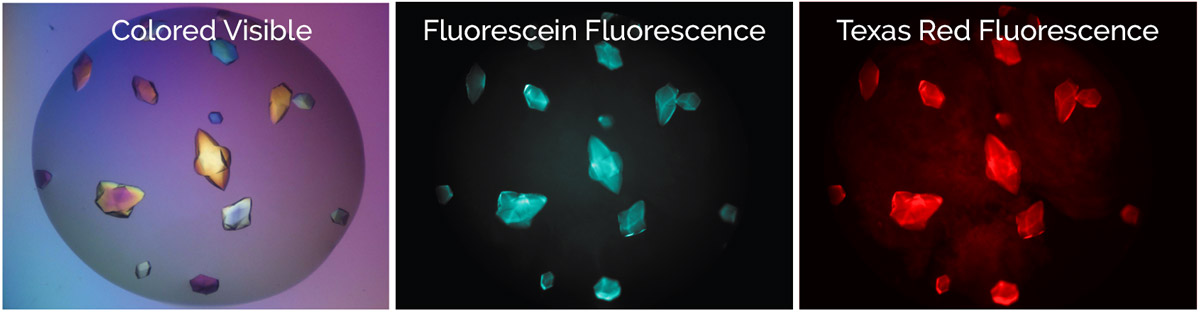
Bovine pancreatic trypsin inhibitor (BPTI) labeled with Fluorescein and trypsin labeled with Texas Red.
Crystals fluoresce at both wavelengths indicating that they are the BPTI-trypsin complex.1
Image Using 3 Different Wavelengths of Your Choice
Choose up to 3 different light wavelengths to image your crystals, including ultraviolet and various visible light wavelengths. Interchangeable filter modules are used to maximize image quality and dye-label compatibility.
Designed for Image Quality
MFI’s optics are optimized for both UV and visible fluorescence imaging to provide high-contrast images. The UV LED and condenser lenses are positioned to maximize the intensity of the UV illumination in order to boost signal strength and produce quality UV images. For visible fluorescence imaging, individual filter cubes with dedicated LED light sources are used to maximize efficiency for each of the chosen wavelengths.
See More with High Magnification Objectives
With two objectives, you can zoom out to see the entire drop or zoom in to see your crystals in greater detail. Both objectives are located on a motorized wheel for quick and easy, automated selection and imaging. The table below provides the optical specifications for the different objectives.
| Objective | Numerical Aperture | Depth of Field (mm) | Field of View (mm) | Pixel Size (µL) |
|---|---|---|---|---|
| 3.3x | 0.11 | 0.1 | 3.7 x 3.0 | 1.1 |
| 6.6x | 0.23 | 0.05 | 1.9 x 1.5 | 0.56 |

Quick + Safe Labeling that Lasts
Simple 0.1% protein labeling protocol
1. Prepare a 5mM stock solution of a succinimidyl ester dye in DMSO
2. Add appropriate amount of dye to protein solution for 0.1% labeling of lysine residues assuming a 1:1 stoichiometric labeling efficiency to amine residues
3. Wait 5 minutes at which point 90% of the dye is bound
There is no need for purification and samples are still fluorescent after 120 days. Protein labeling with fluorophores has been shown to have no known negative impact on crystallization and demonstrated as a useful method for positively identifying protein crystallization hits2,3,4,5
Options to Fit your Needs + Budget
MFI is an option available in our Rock Imager 2 and Rock Imager 360 benchtop imaging systems, or in our Rock Imager 1000 system with automated plate storage.
All these models can automatically image plates in visible light and up to 3 different wavelengths of your choosing.
FAQs
What is multi-fluorescence imaging (MFI) and how is it used in protein crystallization research?
Multi-Fluorescence Imaging (MFI) is an advanced imaging technique that uses optimized UV and visible fluorescence optics to produce high-contrast crystal images. The Rock Imager MFI system features three interchangeable filter modules, enabling imaging at different excitation and emission wavelengths — typically including a 290 nm UV excitation module and two visible fluorescence filter sets for Texas Red and Fluorescein dyes. In protein crystallization research, MFI is used to distinguish protein crystals from salt crystals and to determine whether crystals are formed from a single protein or a protein–protein complex.
How does multi-fluorescence imaging distinguish protein crystals from salt crystals in crystallization experiments?
MFI differentiates protein crystals from salt crystals by detecting fluorescence signals unique to proteins. Using the UV module, MFI captures fluorescence emitted by aromatic amino acids such as tryptophan, which are naturally present in protein crystals. Alternatively, if the protein is pre-labeled with a fluorescent dye, MFI can image the crystallization drop at the dye’s specific excitation and emission wavelengths. In both cases, protein crystals appear fluorescent while salt crystals remain dark, making them easy to distinguish.
Can multi-fluorescence imaging (MFI) differentiate between protein–protein complex crystals and single-protein crystals?
Yes. MFI can distinguish between crystals of a protein–protein complex and those of a single protein. Each protein or subunit is labeled with a different amine-reactive fluorescent dye before crystallization. The crystallization drop is then imaged at the two corresponding wavelengths. Crystals that fluoresce at both wavelengths indicate the presence of a protein–protein complex, while crystals that fluoresce at only one wavelength correspond to single-protein crystals.